Models
The espm.models module implements the model objects that are used to simulate features of the data (e.g. in EDXS it corresponds to spectra).
The models are also used for the creation of the G matrix, which can be later used for the decomposition of data using NMF.
Note
For now espm supports only the modelling of EDS data but we aim at supporting the modelling of EELS data too.
Models base classes
The PhysicalModel is the abstract class from which the EDX model inherits. The ToyModel can be used for testing data analysis algorithms.
- class espm.models.base.PhysicalModel(e_offset, e_size, e_scale, params_dict, db_name='default_xrays.json', E0=200, **kwargs)[source]
Abstract class of the models
- abstract NMF_initialize_W(D)[source]
Function to be called when initializing the NMF optimization. It returns the matrices W. The basic implementation is to return the pseudo-inverse of the matrix G.
- Parameters:
- D
- np.array 2D:
scikit-learn initialization matrix. It should have the shape (n_features, n_components).
- Returns:
- W
- np.array 2D:
The initialized matrix W of shape (n_G, n_components)
- abstract NMF_simplex()[source]
Function to be called when using the simplex constraint in the ESpM-NMF. Its purpose is to return the indices of the matrix W that will be considered to apply the simplex constraint.
Returns :
- indices :
- np.array 1D:
Indices to be considered to apply the simplex constraint
- abstract NMF_update(W=None)[source]
Function to be called when during the NMF optimization. It returns the matrix G, updated if necessary. It should be run in between each W iteration. You do not need to implement it, if you do not need to update the matrix G during the optimization.
- Parameters:
- W
- np.array 2D:
The part of the matrix W that is required to update the matrix G.
- Returns:
- G
- np.array 2D:
The updated matrix G
- build_energy_scale(e_offset, e_size, e_scale)[source]
Build an array corresponding to a linear energy scale.
- Parameters:
- e_offset
- float:
Offset of the energy scale
- e_size
- int:
Number of channels of the detector
- e_scale
- float:
Scale in keV/channel
- Returns:
- energy scale
- np.array 1D:
Linear energy scale based on the given parameters of shape (e_size,)
- class espm.models.base.ToyModel(L: int = 200, C: int = 15, K: int = 3, seed: int | None = None)[source]
Toy model.
Simple model with a random G matrix and phases.
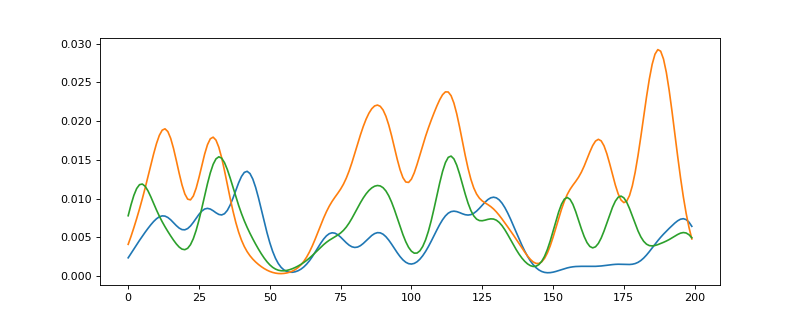
The G matrix is generated with a random number of gaussians per column. The phases are generated using a vector drawn from a Laplacian distribution and multiplied by the G matrix.
- Parameters:
- Lint
Length of the phases.
- Cint
Number of possible components in the phases.
- Kint
Number of phases.
- seedint, optional
Seed for the random number generator.
Examples
>>> from espm.models import ToyModel >>> import matplotlib.pyplot as plt >>> model = ToyModel(L=200, C=15, K=3, seed=0) >>> model.generate_g_matr() >>> model.generate_phases() >>> print(model.G.shape) (200, 15) >>> print(model.phases.shape) (200, 3) >>> plt.plot(model.phases)
EDXS model
The espm.models.edxs module implements the creation of the G matrix that contains a modelisation of the characteristic and continuous X-rays.
This module is a key component of the EDXS data simulation and the EDXS data analysis.
- class espm.models.edxs.EDXS(*args, width_slope=0.01, width_intercept=0.065, custom_init=False, **kwargs)[source]
- NMF_initialize_W(D)[source]
Function to be called when initializing the NMF optimization. It returns the matrices W. The basic implementation is to return the pseudo-inverse of the matrix G.
- Parameters:
- D
- np.array 2D:
scikit-learn initialization matrix. It should have the shape (n_features, n_components).
- Returns:
- W
- np.array 2D:
The initialized matrix W of shape (n_G, n_components)
- NMF_simplex()[source]
Produce the indices of the rows of W on which to perform the simplex constraint.
- generate_g_matr(g_type='bremsstrahlung', *, elements=[], elements_dict={}, **kwargs)[source]
Generate the G matrix. With a complete model the matrix is (e_size,n+2). The first n columns correspond to the sum of X-ray characteristic peaks associated to each shell of the elements. The last 2 columns correspond to a bremsstrahlung model.
The first n columns of G (noted \(\kappa\)), corresponding to the characteristic X-rays, can be calculated using the following formula:
\[\kappa_{k,Z} = \sum_{ij} x_{ij}(Z) \frac{1}{\Delta(\varepsilon_k) \sqrt{2\pi} } e{^{-\frac{1}{2} {\left( \frac{\varepsilon_k - \varepsilon^{ij}(Z)}{\Delta(\varepsilon_k)} \right)}^2}}\]where \(\varepsilon_k\) is the energy of the kth energy channel, \(\varepsilon^{ij}(Z)\) is the energy of the ijth line of the Zth element, \(\Delta(\varepsilon_k)\) is the FWHM of the detector at the energy \(\varepsilon_k\) and \(x_{ij}(Z)\) is the intensity ratio of the ijth line of the Zth element. The last term of the equation is a gaussian function centered at \(\varepsilon^{ij}(Z)\) and with a FWHM of \(\Delta(\varepsilon_k)\).
The last two columns of G (noted \(\beta\)), corresponding to the bremsstrahlung model, can be calculated using the following formula:
\[ \begin{align}\begin{aligned}\beta_{k,n+1} = \frac{\varepsilon_0 - \varepsilon_k}{\varepsilon_0 \varepsilon_k} \times \frac{1 - (\varepsilon_0 - \varepsilon_k)}{\varepsilon_0}\\\beta_{k,n+2} = \frac{(\varepsilon_0 - \varepsilon_k)^2}{\varepsilon^2_0 \varepsilon_k}\end{aligned}\end{align} \]Note that there is no parameter for the bremsstrahlung since it is supposed to be learned with the W matrix (see espm.estimators).
- Parameters:
- g_type
- string:
Options of the edxs model to include in the G matrix. The three possibilities are “identity”, “no_brstlg” and “bremsstrahlung”. G is going to be the identity matrix, only characteristic X-ray peaks or characteristic X-rays plus bremsstrahlung model, respectively.
- elements_dict
- dict:
The keys are chemical elements (atomic number) and the values are cut-off energies. This argument is used to split some of the columns of G into 2 columns. The first column corresponds to characteristic X-rays before the cut-off and second one corresponds to characteristic X-rays before the cut-off. This feature is implemented to enable more accurate absorption correction.
- elements
- list:
List of modeled chemical elements. The list can be populated either with atomic numbers or chemical symbols, e.g. “Fe” or 26.
- Returns:
- g matrix
- np.array 2D:
matrix of the edx model.
Notes
See our paper about the espm package [TPH+23] for more information about the equations of this model.
- generate_phases(phases_parameters)[source]
Generate a series of spectra from list of phase parameters.
- Parameters:
- phases_parameters
- list:
List of dicts containing the phase parameters.
- Returns:
- phase_array
- np.array 2D:
Array which lines correspond to the modelled phases in the input list.
Notes
The absorption correction is done using the average composition of the phases. The same correction is used for each phase.
- generate_spectrum(b0=0, b1=0, scale=1.0, abs_elts_dict={}, *, elements_dict={})[source]
Generate a spectrum from bremsstrahlung parameters and a chemical composition. The modelling is done using the same formula as for the generate_g_matr function.
- Parameters:
- b0
- float:
First bremsstrahlung parameter.
- b1
- float:
Second bremsstrahlung parameter.
- scale
- float:
Scale factor to apply to the bremsstrahlung. Feature to be removed, scaling should be done with b0 and b1.
- abs_elts_dict
- dict:
Dictionnary of elements and associated concentrations. This dictionnary is used to calculate the contribution of the absorption in the spectrum.
- elements_dict
- dict:
Dictionnary of elements and associated concentrations describing the chemical composition of the modeled sample.
- Returns
- ——-
- spectrum
- np.array 1D:
Output edx spectrum corresponding to the input chemical composition.
Notes
Check EDXS_function for details about the bremsstrahlung model.
Examples
>>> import matplotlib.pyplot as plt >>> from espm.models.edxs import EDXS >>> from espm.conf import DEFAULT_EDXS_PARAMS >>> b0, b1 = 5.5367e-5, 0.00192181 >>> elts_dict = {"Si" : 1.0,"Ca" : 1.0,"O" : 3.0,"C" : 0.3} >>> model = EDXS(**DEFAULT_EDXS_PARAMS) >>> spectrum = model.generate_spectrum(b0,b1, elements_dict = elts_dict) >>> plt.plot(spectrum)

EDXS functions
The espm.models.EDXS_function module implements misceallenous functions required for the :mod: espm.models.edxs.
This module include bremsstrahlung modelling, database reading and dicts and list manipulation functions.
- espm.models.EDXS_function.G_bremsstrahlung(x, E0, params_dict, *, elements_dict={})[source]
Computes the two-parts continuum X-rays for the G matrix. The two parts of the bremsstrahlung are constructed separately so that its parameters can fitted to data. Absorption and detection are multiplied to each part.
- Parameters:
- x
- np.array 1D:
Energy scale.
- params_dict
- dict:
Dictionnary containing the absorption and detection parameters.
- elements_dict
- dict:
Composition of the studied sample. It is required for absorption calculation.
- Returns:
- continuum_xrays
- np.array 2D:
Two parts continuum X-rays model with shape (energy scale size, 2).
- espm.models.EDXS_function.chapman_bremsstrahlung(x, b0, b1, b2)[source]
Calculate the bremsstrahlung as parametrized by chapman et al.
- Parameters:
- x
- np.array 1D:
Energy scale.
- b0
- float:
First parameter, corresponding to the inverse of the energy.
- b1
- float:
Second parameter, corresponding to the energy.
- b2
- float:
Third parameter, corresponding to the square of the energy.
- Returns:
- bremsstrahlung
- np.array 1D:
Bremsstrahlung model
Notes
For details see [CNC84]
- espm.models.EDXS_function.continuum_xrays(x, params_dict={}, b0=0, b1=0, E0=200, *, elements_dict={'Si': 1.0})[source]
Computes the continuum X-rays, i.e. the bremsstrahlung multiplied by the absorption and the detection efficiency.
- Parameters:
- x
- np.array 1D:
Energy scale.
- params_dict
- dict:
Dictionnary containing the absorption and detection parameters.
- b0
- float:
First parameter.
- b1
- float:
Second parameter.
- E0
- float:
Energy of the incident beam in keV.
- elements_dict
- dict:
Composition of the studied sample. It is required for absorption calculation.
- Returns:
- continuum_xrays
- np.array 1D:
Continuum X-rays model.
Notes
When an empty dict is provided. The continuum X-rays are set to 0. This is useful to perform calculations without the bremsstrahlung for example.
For an example structure of the params_dict parameter, check the DEFAULT_EDXS_PARAMS espm.conf.
For a custom detection efficiency, check the spectrum fit notebook.
- espm.models.EDXS_function.elts_dict_from_dict_list(dict_list)[source]
Create a single dictionnary of the elemental concentration from a list of dictionnary containing chemical compositions. The new dictionnary corresponds to the average chemical composition of the list.
- Parameters:
- dict_list
- list [dict]:
list of chemical composition dictionnary
- Returns:
- elements_dictionnary
- dict:
Dictionnary containing the concentration associated to each chemical elements contained in the input.
Notes
This function should maybe move to another part of espm.
Examples
>>> import numpy as np >>> from espm.models.EDXS_function import elts_dict_from_dict_list >>> dicts = [{"Si" : 1.0},{"Ca" : 1.0},{"O" : 1.0},{"C" : 1.0}] >>> elts_dict_from_dict_list(dicts) {"Si" : 0.25, "Ca" : 0.25, "O" : 0.25, "C" : 0.25}
- espm.models.EDXS_function.elts_list_from_dict_list(dict_list)[source]
Create a list of elements from a list of dictionnary containing chemical compositions. The list contains all the elements contained in the input.
- Parameters:
- dict_list
- list [dict]:
list of chemical composition dictionnary
- Returns:
- elements_list
- list:
List of elements as atomic number or symbol.
Examples
>>> import numpy as np >>> from espm.models.EDXS_function import elts_list_from_dict_list >>> dicts = [{"Si" : 1.0, "C" : 0.5, "O" : 0.5},{"Ca" : 1.0, "K" : 2.0},{"O" : 1.0, "Si" : 0.5},{"C" : 1.0}] >>> elts_list_from_dict_list(dicts) ["Si","Ca","O","C","K"]
- espm.models.EDXS_function.gaussian(x, mu, sigma)[source]
Calculate the gaussian function according to the following formula:
\[f(x) = \frac{1}{\sigma \sqrt{2 \pi}} e^{-\frac{(x-\mu)^2}{2 \sigma^2}}\]- Parameters:
- xnp.array 1D
Energy scale.
- mufloat
Mean of the gaussian.
- sigmafloat
Standard deviation of the gaussian.
- Returns:
- gaussiannp.array 1D
Gaussian distribution on x, with mean mu and standard deviation sigma.
- espm.models.EDXS_function.lifshin_bremsstrahlung(x, b0, b1, E0=200)[source]
Calculate the custom parametrized bremsstrahlung inspired by the model of L.Lifshin.
The model is designed to have the same definition set as the original model of L.Lifshin with \(b_0 > 0\) and \(b_1 > 0\).
The model is defined as follows:
\[f(\varepsilon) = b_0 \frac{\varepsilon_0 - \varepsilon}{\varepsilon_0 \varepsilon} \left(1 - \frac{\varepsilon_0 - \varepsilon}{\varepsilon_0}\right) + b_1 \frac{\left( \varepsilon_0 - \varepsilon \right) ^2}{\varepsilon_0^2 \varepsilon}\]where \(\varepsilon_0\) is the energy of the incident beam, \(\varepsilon\) is the energy of the emitted photon and \(b_0\) and \(b_1\) are the parameters of the model.
- Parameters:
- x
- np.array 1D:
Energy scale.
- b0
- float:
First parameter.
- b1
- float:
Second parameter.
- E0
- float:
Energy of the incident beam in keV.
- Returns:
- bremsstrahlung
- np.array 1D:
Bremsstrahlung model
Notes
For details see L.Lifshin, Ottawa, Proc.9.Ann.Conf.Microbeam Analysis Soc. 53. (1974)
- espm.models.EDXS_function.lifshin_bremsstrahlung_b0(x, b0, E0=200)[source]
Calculate the first part of our bremsstrahlung model.
- espm.models.EDXS_function.lifshin_bremsstrahlung_b1(x, b1, E0=200)[source]
Calculate the second part of our bremsstrahlung model.
- espm.models.EDXS_function.read_compact_db(elt, db_dict)[source]
Read the energy and cross section of each line of a chemical element for compact databases.
- Parameters:
- elt
- string:
Atomic number.
- db_dict
- dict:
Dictionnary extracted from the json database containing the emission lines and their energies.
- Returns:
- energies
- list float:
List of energies associated to each emission line of the given element
- cross-sections
- list float:
List of emission cross-sections of each line of the given element
- espm.models.EDXS_function.read_lines_db(elt, db_dict)[source]
Read the energy and cross section of each line of a chemical element for explicit databases.
- Parameters:
- elt
- string:
Atomic number.
- db_dict
- dict:
Dictionnary extracted from the json database containing the emission lines and their energies.
- Returns:
- energies
- list float:
List of energies associated to each emission line of the given element
- cross-sections
- list float:
List of emission cross-sections of each line of the given element
- espm.models.EDXS_function.shelf(x, height, length)[source]
Calculate the shelf contribution of the EDXS Silicon Drift Detector (SDD) detector.
- Parameters:
- x
- np.array 1D:
Energy scale.
- height
- float:
Height in intensity of shelf contribution of the detector.
- length
- float:
Length in energy of shelf contribution of the detector.
- Returns:
- shelf
- np.array 1D:
SDD shelf model.
Notes
For details see [SP09]
Absorption functions
The espm.models.absorption_edxs module implements the functions to calculate the contribution of absorption in edx spectra.
- espm.models.absorption_edxs.absorption_coefficient(x, atomic_fraction=False, *, elements_dict={'Si': 1.0})[source]
Calculate the mass-absorption coefficent of a given composition at a certain energy or over a certain energy range.
- Parameters:
- x
- np.array 1D:
Energy value or energy scale over which the mass-absorption coefficient is calculated.
- atomic_fraction
- boolean:
If True, the atomic concentrations of elements_dict are converted in atomic weight fractions.
- elements_dict
- dict:
Dictionnary with chemical elements as keys and atomic weight fractions (or atomic concentrations) as values.
- Returns:
- mass-absorption coefficient
- np.array 1D:
Calculated value of the mass-absorption coefficient on all the energy range.
Notes
The mass-absorption coefficients are calculated using the database of hyperspy [dlPPF+22].
- espm.models.absorption_edxs.absorption_correction(x, thickness=1e-05, toa=90, density=None, atomic_fraction=False, *, elements_dict={'Si': 1.0}, **kwargs)[source]
Calculate the contribution of the absorption on the EDX spectrum for a thin slab of material with a given composition at a certain energy or over a certain energy range. The absorption correction is calculated using the following formula :
\[\frac{1 - \exp(-\chi)}{\chi} \]where \(\chi = \mu \rho t / \sin(\theta)\), \(\mu\) is the mass-absorption coefficient, \(\rho\) is the density of the material, \(t\) is the thickness of the sample and \(\theta\) is the average take-off angle of the x-rays travelling from the sample to the x-ray detectors.
- Parameters:
- x
- np.array 1D:
Energy value or energy scale over which the mass-absorption coefficient is calculated.
- thickness
- float:
Thickness of the material slab in meter. If the thickness is set to 0.0, the function will return 1.0.
- toa
- float:
Take-off angle in degrees of the x-rays travelling from the sample to the x-ray detectors.
- density
- float:
Density of the material in g/m3 (to be checked). If None, an approximation of the density based on the atomic numbers of the elements of the materials is calculated.
- atomic_fraction
- boolean:
If True, the atomic concentrations of elements_dict are converted in atomic weight fractions.
- elements_dict
- dict:
Dictionnary with chemical elements as keys and atomic weight fractions (or atomic concentrations) as values.
- Returns:
- absorption correction
- np.array 1D:
Calculated value of the absorption correction on all the energy range.
- espm.models.absorption_edxs.absorption_mass_thickness(x, mass_thickness, toa=90, atomic_fraction=True, *, elements_dict={'Si': 1.0})[source]
Calculate the contribution of the absorption of a mass-thickness map.
- Parameters:
- x
- np.array 1D:
Energy value or energy scale over which the mass-absorption coefficient is calculated.
- mass_thickness:
- np.array 2D:
Mass-thickness map. See the notes for details.
- toa
- float:
Take-off angle in degrees of the x-rays travelling from the sample to the x-ray detectors.
- atomic_fraction
- boolean:
If True, the atomic concentrations of elements_dict are converted in atomic weight fractions.
- elements_dict
- dict:
Dictionnary with chemical elements as keys and atomic weight fractions (or atomic concentrations) as values.
- Returns:
- absorption correction
- np.array 3D:
Calculated value of the absorption correction on all the energy range for all pixels of the mass-thickness map.
Notes
The mass-thickness map can be extracted from an EDX spectrum image by calculating the concentration of an element for both K and L lines. Since there should be only one concentration for both lines, it is possible to determine the contribution of the absorption for each pixel and thus obtain the mass-thickness map.
- espm.models.absorption_edxs.det_efficiency(x, det_dict)[source]
Simulate the detection efficiency of an EDX detector. The absorption wihtin one layer of the detector is calculated according to the following equation, corresponding to the Beer-Lambert law:
\[d_k = \exp\left( - \mu(\varepsilon_k)\rho t \right)\]where \(d_k\) is the detection efficiency at the energy \(\varepsilon_k\), \(\mu(\varepsilon_k)\) is the mass-absorption coefficient at the energy \(\varepsilon_k\), \(\rho\) is the density of the material, and \(t\) is the thickness of the material.
In a detection layer of the detector, an absobed photon counts as a detected photon so that the detection efficiency is equal to 1 minus the absorption. The detection efficiency is calculated as the product of the absorption of each layer of the detector.
- Parameters:
- x
- np.array 1D:
Energy value or energy scale over which the mass-absorption coefficient is calculated.
- det_dict
- dict:
See Examples for an example of the structure of the dict. One of the layer of the input dictionnary should be named “detection” to model a active layer of the detector.
- Returns:
- efficiency
- np.array 1D:
Calculated detection efficiency based on the input layers model.
Examples
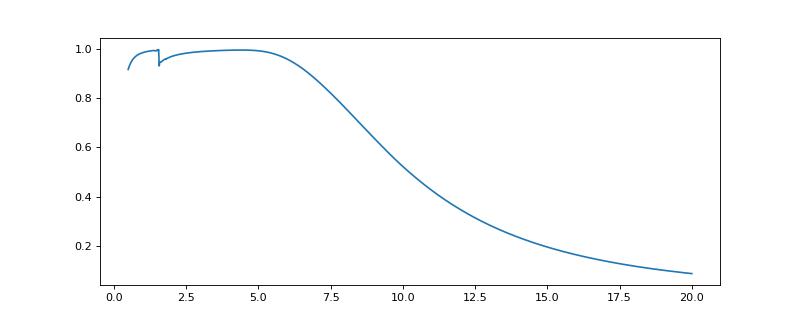
>>> from espm.models.absorption_edxs import det_efficiency >>> import matplotlib.pyplot as plt >>> import numpy as np >>> dict = {"detection" : {"thickness" : 10e-3, "density" : 2.3, "atomic_fraction" : False, "elements_dict" : {"Si" : 1.0}}, >>> "dead_layer" : {"thickness" : 50e-7, "density" : 2.7, "atomic_fraction" : False, "elements_dict" : {"Al" : 1.0}}} >>> x = np.linspace(0.5,20,num = 1000) >>> efficiency = det_efficiency(x,dict) >>> plt.plot(x,efficiency)
- espm.models.absorption_edxs.det_efficiency_from_curve(x, filename, kind='cubic')[source]
Interpolate a detection efficiency curve that is stored in ~/espm/tables.
- Parameters:
- x
- np.array 1D:
Energy value or energy scale over which the mass-absorption coefficient is calculated.
- filename:
- string:
Name of the file of the detection efficiency.
- kind
- string:
Polynomial order of the interpolation.
- Returns:
- Detection efficiency
- np.array 1D:
Interpolated detection efficiency.
- espm.models.absorption_edxs.det_efficiency_layer(x, thickness=1e-05, density=None, atomic_fraction=False, *, elements_dict={'Si': 1.0})[source]
Calculate the proportion of absorbed X-rays for one virtual layer of the detector.
- Parameters:
- x
- np.array 1D:
Energy value or energy scale over which the mass-absorption coefficient is calculated.
- thickness
- float:
Thickness of the material slab in meter. If the thickness is set to 0.0, the function will return 1.0.
- density
- float:
Density of the material in g/m3 (to be checked). If None, an approximation of the density based on the atomic numbers of the elements of the materials is calculated.
- atomic_fraction
- boolean:
If True, the atomic concentrations of elements_dict are converted in atomic weight fractions.
- elements_dict
- dict:
Dictionnary with chemical elements as keys and atomic weight fractions (or atomic concentrations) as values.
- Returns:
- absorption
- np.array 1D:
Calculated absorption of one layer of the detector.
Notes
We assume that the X-rays arrive perpendicular to the detector.
- espm.models.generate_EDXS_phases.generate_brem_params(seed)[source]
Generate random parameters for the Bremsstrahlung model with a scaling factor that somewhat resembles the real data.
- Parameters:
- seedint
Seed for the random number generator.
- Returns:
- dict
Dictionary containing the parameters for the Bremsstrahlung model.
- espm.models.generate_EDXS_phases.generate_elts_dict(seed, nb_elements=3)[source]
Generate a random dictionary of elements and their relative abundance. The elements are limited to the range 6-82.
- Parameters:
- seedint
Seed for the random number generator.
- nb_elementsint, optional
Number of elements to generate. The default is 3.
- Returns:
- dict
Dictionary containing the elements and their relative abundance.
- espm.models.generate_EDXS_phases.generate_modular_phases(elts_dicts=3, brstlg_pars=None, scales=None, model_params=None, seed=0)[source]
Generate an array of phases of the EDXS model with set model parameters, elemental compositions and bremsstrahlung parameters.
- Parameters:
- elts_dictsint or list, optional
Number of phases to generate or list of dictionnaries containing the chemical compositions. The default is 3.
- brstlg_parslist, optional
List of dictionnaries containing the bremsstrahlung parameters. If the parameters are not set, they are generated randomly.
- scaleslist, optional
List of scaling factors for the phases. If the scales are not set, they are set to 1.0.
- model_paramsdict, optional
Dictionary containing the model parameters. If the parameters are not set, they are set to the default values. See the config file for the default values.
- seedint, optional
Seed for the random number generator. The default is 0.
- Returns:
- np.array
Array of phases of the EDXS model.
Examples
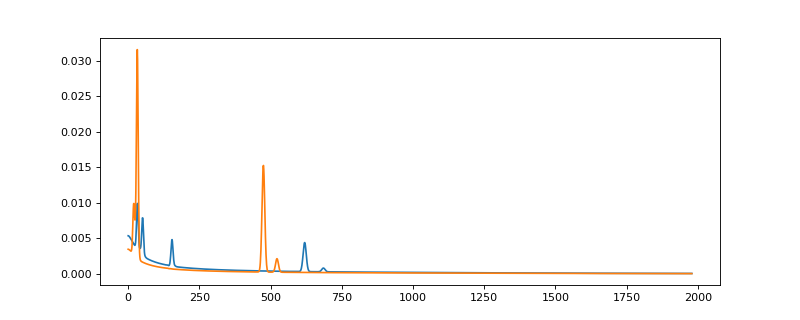
>>> from espm.models.generate_EDXS_phases import generate_modular_phases >>> import matplotlib.pyplot as plt >>> phases = generate_modular_phases(elts_dicts = [{"8": 0.5, "14": 0.2, "26": 0.3}, {"8" : 0.5, "23" : 0.3, "7" : 0.2}], ... brstlg_pars = [{"b0" : 0.03, "b1" : 0.01}, {"b0" : 0.002, "b1" : 0.0025}]) >>> plt.plot(phases[0]) >>> plt.plot(phases[1])
- espm.models.generate_EDXS_phases.generate_random_phases(n_phases=3, seed=0)[source]
Generate a list of phases of the EDXS model with random elemental compositions and Bremsstrahlung parameters. The model parameters are set to the default values.
- Parameters:
- n_phasesint, optional
Number of phases to generate. The default is 3.
- seedint, optional
Seed for the random number generator. The default is 0.
- Returns:
- np.array
Array of phases of the EDXS model.
- espm.models.generate_EDXS_phases.unique_elts(dict_list)[source]
Generate a list of unique chemical elements from a list of chemical elements dictionnaries.
- Parameters:
- dict_listlist
List of dictionnaries containing the elements and their relative abundance.
- Returns:
- list
List of unique chemical elements.
Examples
>>> unique_elts([{"elements_dict" : {"18" : 0.2, "8" : 0.5, "12" : 0.3}}, {"elements_dict" : {"8" : 0.5, "26" : 0.3, "14" : 0.2}}]) ['18', '8', '12', '26', '14']